Running compute jobs#
Now we’re connected to Picotte—but Picotte, like we saw in
the Introduction, is composed of many computers. Connecting to
picottelogin.urcf.drexel.edu just connects us to one of them, the so-called
“login node” (“node” just means one of the computers that make up the cluster).
But Picotte has nearly 100 nodes in it—how do we access the others?
The login node is a gateway to where the real work happens—the other nodes of the cluster, which are called “compute nodes”. You can think of this like a busy medical office. The login node is the waiting room, and the compute nodes are the examination rooms (where the work actually happens).
To access the compute nodes we need to submit a request to the scheduler. The scheduler accepts requests for the cluster to do some work (called jobs), keeps track of all pending jobs, and assigns them to compute nodes to run as capacity becomes available. The scheduler helps ensure that the cluster’s resources are used fairly by balancing competing requests from different users and research groups[1].
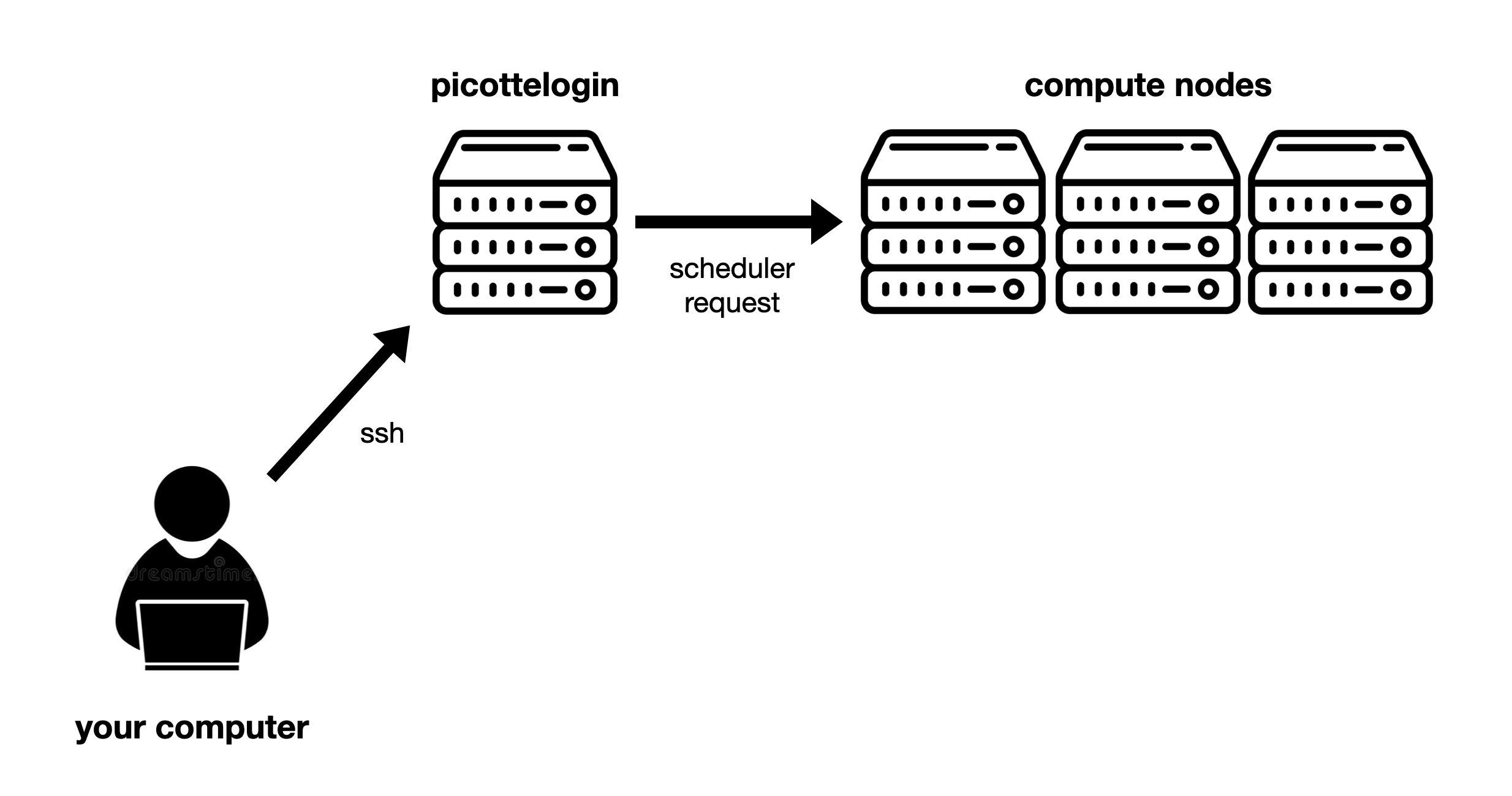
Fig. 7 Overview of Picotte access: From your computer, you access the login node using SSH. From there, you access the compute nodes by submitting requests to the scheduler.#
Our first job#
The scheduler used by Picotte is called SLURM. SLURM has many commands, we’ll learn a few of the most important today.
SLURM commands generally start with the letter s.
Let’s run our first job, using the srun command[2] as follows:
srun --partition=def-sm --account=workshopprj Rscript -e 'summary(mtcars)'
The Rscript command runs a program using the R
language for statistical computing. In this case,
we’re computing summary statistics on the built-in mtcars dataset[3].
You should see something like this:
[jjp366@picotte001 ~]$ srun --partition=def-sm --account=workshopprj Rscript -e 'summary(mtcars)'
Microsoft R Open 4.0.2
The enhanced R distribution from Microsoft
Microsoft packages Copyright (C) 2020 Microsoft Corporation
Using the Intel MKL for parallel mathematical computing (using 1 cores).
Default CRAN mirror snapshot taken on 2020-07-16.
See: https://mran.microsoft.com/.
mpg cyl disp hp
Min. :10.40 Min. :4.000 Min. : 71.1 Min. : 52.0
1st Qu.:15.43 1st Qu.:4.000 1st Qu.:120.8 1st Qu.: 96.5
Median :19.20 Median :6.000 Median :196.3 Median :123.0
Mean :20.09 Mean :6.188 Mean :230.7 Mean :146.7
3rd Qu.:22.80 3rd Qu.:8.000 3rd Qu.:326.0 3rd Qu.:180.0
Max. :33.90 Max. :8.000 Max. :472.0 Max. :335.0
drat wt qsec vs
Min. :2.760 Min. :1.513 Min. :14.50 Min. :0.0000
1st Qu.:3.080 1st Qu.:2.581 1st Qu.:16.89 1st Qu.:0.0000
Median :3.695 Median :3.325 Median :17.71 Median :0.0000
Mean :3.597 Mean :3.217 Mean :17.85 Mean :0.4375
3rd Qu.:3.920 3rd Qu.:3.610 3rd Qu.:18.90 3rd Qu.:1.0000
Max. :4.930 Max. :5.424 Max. :22.90 Max. :1.0000
am gear carb
Min. :0.0000 Min. :3.000 Min. :1.000
1st Qu.:0.0000 1st Qu.:3.000 1st Qu.:2.000
Median :0.0000 Median :4.000 Median :2.000
Mean :0.4062 Mean :3.688 Mean :2.812
3rd Qu.:1.0000 3rd Qu.:4.000 3rd Qu.:4.000
Max. :1.0000 Max. :5.000 Max. :8.000
[jjp366@picotte001 ~]$
The output includes statistics (e.g. mean, median) for each of the variables on car performance: miles per gallon, horsepower, etc. The details of the computation and output don’t really matter here, we’re just using it as a simple example of running a job on the cluster.
Let’s trace what happened: the job was queued, SLURM allocated resources on a compute node, the R script ran there, the output was printed back to your terminal, and then you were returned to the login node. The key takeaway is that SLURM handled scheduling the work onto a compute node transparently—you didn’t have to know or care which node it ran on.
The cluster just did real work for us—it ran an R script on a compute node and returned the results. The command we ran here was simple, but the same approach works for anything your research requires: a Python script you wrote, open-source bioinformatics tools, commercial structural analysis software—the sky’s the limit. We have a lot of research software pre-installed on Picotte for you to use.
Let’s break down exactly what each of the arguments in the command above means and why we need them:
srun
The base command: this launches a single job on a compute node.
Rscript -e 'summary(mtcars)'
the main argument to
srunis the command to run. Here, we’re runningRscript -e 'summary(mtcars)', which executes an R expression that summarizes the built-inmtcarsdataset.
--partition=def-sm and --account=workshopprj
These options specify the account and partition the job should run under. These concepts merit more explanation in the next section.
Accounts and partitions#
Our first job submission immediately showed us two of the most important SLURM
concepts: accounts (the --account argument) and partitions (the --partition
argument).
Accounts#
Confusingly, this is not your Picotte user account. SLURM has a separate
concept of “bank accounts” which is what’s referred to here. You’ll also see
these referred to as “SLURM accounts” or “projects” (and they typically end in
prj).
Every job submitted to the scheduler has to be associated with a single SLURM account, which will be billed for the job’s usage.
Typically, each research group has two projects:
groupnameprjfor paid, priority tier jobsgroupnamefreeprjfor free-tier jobs
For this workshop, we’re all just using workshopprj which can only run
free-tier jobs. Picotte has two “tiers” of access, free and priority. You can
read more about Picotte usage rates and tiers on our documentation
site.
You can see all the SLURM accounts you have access to using the command:
sacctmgr show user $USER withassoc
Partitions#
A “partition” is a group of compute nodes. Partitions are also sometimes called “queues” or “job queues”, because jobs are scheduled separately on different partitions, so each is sort of has it’s own “queue” of jobs waiting to run.
Partitions are typically groups of nodes with similar hardware. For example, the
gpu partition includes all Picotte nodes with GPUs, and the bm partition
(“big memory”) has nodes with much more memory than the typical compute node.
Partitions also can have different access policies, resource limits, and billing. For example:
def-smis the standard free-tier partition. All free-tier projects have access, and jobs run here are not billed for, but this partition has small resource limits.defis the standard priority-tier partition. Only funded projects have access, and jobs must be paid for, but the resource limits ofdef-smare removed.
All jobs must specify a partition to run in. Here we used the def-sm
partition, which is for standard free-tier jobs. You can read more about all the
available partitions
here.
Picotte is a shared resource
Please be considerate of others when submitting jobs. Remember that Picotte is a shared resource. Don’t request resources you don’t plan on actually using.
Don’t run computations on the login node
When using Picotte, you shouldn’t run heavy computations on the login node, because the login node is shared between everyone who logs into Picotte, so your computations will interfere with other people’s login processes. Once in a job, you can do whatever you want, because the resources assigned to your job (CPUs, memory, etc.) are given to it exclusively, so you won’t interfere with anyone else.
